Python source code: plot_granulo.py
from scipy import ndimage
import matplotlib.pyplot as plt
def disk_structure(n):
struct = np.zeros((2 * n + 1, 2 * n + 1))
x, y = np.indices((2 * n + 1, 2 * n + 1))
mask = (x - n)**2 + (y - n)**2 <= n**2
struct[mask] = 1
return struct.astype(np.bool)
def granulometry(data, sizes=None):
s = max(data.shape)
if sizes == None:
sizes = range(1, s/2, 2)
granulo = [ndimage.binary_opening(data, \
structure=disk_structure(n)).sum() for n in sizes]
return granulo
np.random.seed(1)
n = 10
l = 256
im = np.zeros((l, l))
points = l*np.random.random((2, n**2))
im[(points[0]).astype(np.int), (points[1]).astype(np.int)] = 1
im = ndimage.gaussian_filter(im, sigma=l/(4.*n))
mask = im > im.mean()
granulo = granulometry(mask, sizes=np.arange(2, 19, 4))
plt.figure(figsize=(6, 2.2))
plt.subplot(121)
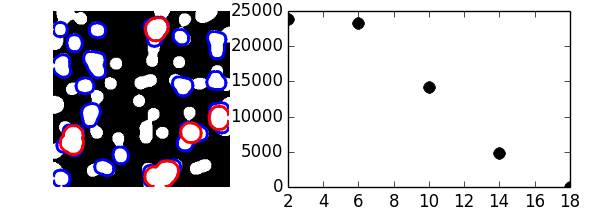
plt.imshow(mask, cmap=plt.cm.gray)
opened = ndimage.binary_opening(mask, structure=disk_structure(10))
opened_more = ndimage.binary_opening(mask, structure=disk_structure(14))
plt.contour(opened, [0.5], colors='b', linewidths=2)
plt.contour(opened_more, [0.5], colors='r', linewidths=2)
plt.axis('off')
plt.subplot(122)
plt.plot(np.arange(2, 19, 4), granulo, 'ok', ms=8)
plt.subplots_adjust(wspace=0.02, hspace=0.15, top=0.95, bottom=0.15, left=0, right=0.95)
plt.show()